Difference between revisions of "AraP"
| Line 31: | Line 31: | ||
__TOC__ | __TOC__ | ||
| − | <br/><br/> | + | <br/><br/><br/><br/><br/><br/> |
=The gene= | =The gene= | ||
Revision as of 14:36, 22 June 2009
- Description: L-arabinose transport (integral membrane protein)
| Gene name | araP |
| Synonyms | yseD |
| Essential | no |
| Product | L-arabinose transport (integral membrane protein)) |
| Function | uptake of arabinose |
| Metabolic function and regulation of this protein in SubtiPathways: Sugar catabolism | |
| MW, pI | 34 kDa, 9.69 |
| Gene length, protein length | 939 bp, 313 aa |
| Immediate neighbours | araQ, araN |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 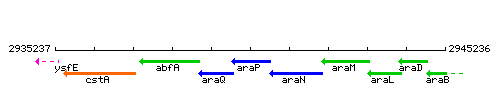
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU28740
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: MalFG subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cell membrane (according to Swiss-Prot)
Database entries
- Structure:
- Swiss prot entry: P94529
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Additional information: the mRNA is very stable (half-life > 15 min) PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
José Manuel Inácio, Carla Costa, Isabel de Sá-Nogueira
Distinct molecular mechanisms involved in carbon catabolite repression of the arabinose regulon in Bacillus subtilis.
Microbiology (Reading): 2003, 149(Pt 9);2345-2355
[PubMed:12949161]
[WorldCat.org]
[DOI]
(P p)
G Hambraeus, C von Wachenfeldt, L Hederstedt
Genome-wide survey of mRNA half-lives in Bacillus subtilis identifies extremely stable mRNAs.
Mol Genet Genomics: 2003, 269(5);706-14
[PubMed:12884008]
[WorldCat.org]
[DOI]
(P p)
L J Mota, P Tavares, I Sá-Nogueira
Mode of action of AraR, the key regulator of L-arabinose metabolism in Bacillus subtilis.
Mol Microbiol: 1999, 33(3);476-89
[PubMed:10417639]
[WorldCat.org]
[DOI]
(P p)
Y Quentin, G Fichant, F Denizot
Inventory, assembly and analysis of Bacillus subtilis ABC transport systems.
J Mol Biol: 1999, 287(3);467-84
[PubMed:10092453]
[WorldCat.org]
[DOI]
(P p)
Isabel S-Nogueira, Teresa V Nogueira, Snia Soares, Hermnia de Lencastre
The Bacillus subtilis L-arabinose (ara) operon: nucleotide sequence, genetic organization and expression.
Microbiology (Reading): 1997, 143 ( Pt 3);957-969
[PubMed:9084180]
[WorldCat.org]
[DOI]
(P p)