Difference between revisions of "Apt"
m (Reverted edits by 134.76.70.252 (talk) to last revision by 134.76.38.147) |
|||
| Line 125: | Line 125: | ||
* '''Additional information:''' | * '''Additional information:''' | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 1738 {{PubMed|21395229}} | ||
| + | |||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 1189 {{PubMed|21395229}} | ||
| + | |||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 1706 {{PubMed|21395229}} | ||
=Biological materials = | =Biological materials = | ||
| − | |||
* '''Mutant:''' | * '''Mutant:''' | ||
Revision as of 13:53, 17 April 2014
- Description: adenine phosphoribosyltransferase, universally conserved protein
| Gene name | apt |
| Synonyms | |
| Essential | no |
| Product | adenine phosphoribosyltransferase |
| Function | purine salvage and interconversion |
| Gene expression levels in SubtiExpress: apt | |
| Metabolic function and regulation of this protein in SubtiPathways: apt | |
| MW, pI | 18 kDa, 4.701 |
| Gene length, protein length | 510 bp, 170 aa |
| Immediate neighbours | relA, yrvE |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 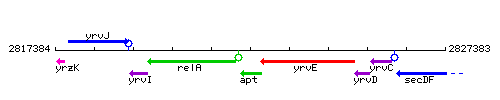
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed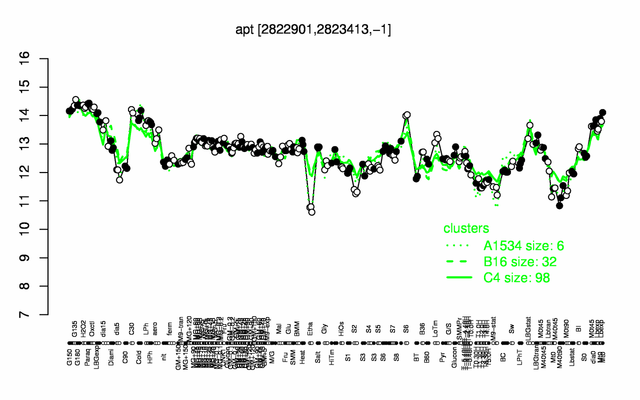
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of nucleotides, universally conserved proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU27610
Phenotypes of a mutant
Database entries
- BsubCyc: BSU27610
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: AMP + diphosphate = adenine + 5-phospho-alpha-D-ribose 1-diphosphate (according to Swiss-Prot)
- Protein family: purine/pyrimidine phosphoribosyltransferase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- phosphorylated on Arg-85 PubMed
- Cofactor(s):
- Effectors of protein activity:
Database entries
- BsubCyc: BSU27610
- Structure: 2DY0 (from Escherichia coli k12, 56% identity, 66% similarity)
- UniProt: O34443
- KEGG entry: [3]
- E.C. number: 2.4.2.7
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 1738 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 1189 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 1706 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Alexander K W Elsholz, Kürsad Turgay, Stephan Michalik, Bernd Hessling, Katrin Gronau, Dan Oertel, Ulrike Mäder, Jörg Bernhardt, Dörte Becher, Michael Hecker, Ulf Gerth
Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis.
Proc Natl Acad Sci U S A: 2012, 109(19);7451-6
[PubMed:22517742]
[WorldCat.org]
[DOI]
(I p)
M Nickel, G Homuth, C Böhnisch, U Mäder, T Schweder
Cold induction of the Bacillus subtilis bkd operon is mediated by increased mRNA stability.
Mol Genet Genomics: 2004, 272(1);98-107
[PubMed:15241682]
[WorldCat.org]
[DOI]
(P p)
H H Saxild, P Nygaard
Genetic and physiological characterization of Bacillus subtilis mutants resistant to purine analogs.
J Bacteriol: 1987, 169(7);2977-83
[PubMed:3110131]
[WorldCat.org]
[DOI]
(P p)