Difference between revisions of "AccD"
| Line 113: | Line 113: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' ''[[accD]]-[[accA]]'' | + | * '''Operon:''' ''[[accD]]-[[accA]]'' {{PubMed|22383849}} |
* '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=accD_2988693_2989565_-1 accD] {{PubMed|22383849}} | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=accD_2988693_2989565_-1 accD] {{PubMed|22383849}} | ||
| Line 149: | Line 149: | ||
<pubmed> 15952903 </pubmed> | <pubmed> 15952903 </pubmed> | ||
==Original Publications== | ==Original Publications== | ||
| − | <pubmed>15066985, 12663926,14651647,16479537 22517742</pubmed> | + | <pubmed>15066985, 12663926,14651647,16479537 22517742 22383849</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 14:08, 21 April 2012
- Description: acetyl-CoA carboxylase (beta subunit)
| Gene name | accD |
| Synonyms | yttI |
| Essential | yes PubMed |
| Product | acetyl-CoA carboxylase (beta subunit)) |
| Function | production of malonyl-CoA, the substrate for fatty acid biosynthesis |
| Interactions involving this protein in SubtInteract: AccD | |
| Metabolic function and regulation of this protein in SubtiPathways: Lipid synthesis | |
| MW, pI | 28 kDa, 5.344 |
| Gene length, protein length | 786 bp, 262 aa |
| Immediate neighbours | accA, ytsJ |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 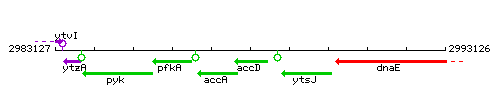
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed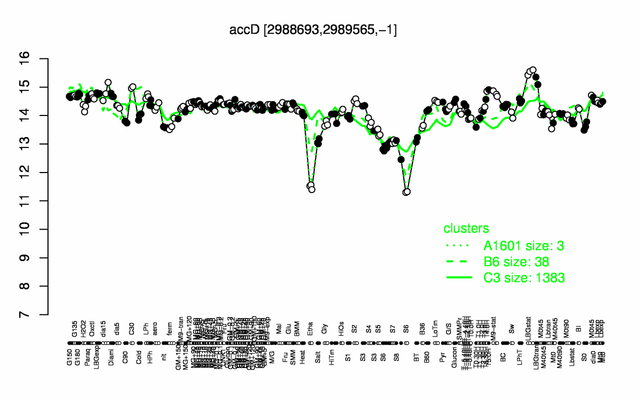
| |
Contents
Categories containing this gene/protein
biosynthesis of lipids, essential genes, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU29210
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- phosphorylated on Arg-205 PubMed
- Cofactor(s):
- Effectors of protein activity:
- Localization: Membrane-proximal (Spotty) PubMed Membrane-proximal (Spotty) PubMed
Database entries
- UniProt: C0SP93
- KEGG entry: [3]
- E.C. number: 6.4.1.2
Additional information
Expression and regulation
- Sigma factor:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Stephen W White, Jie Zheng, Yong-Mei Zhang, Rock
The structural biology of type II fatty acid biosynthesis.
Annu Rev Biochem: 2005, 74;791-831
[PubMed:15952903]
[WorldCat.org]
[DOI]
(P p)
Original Publications
Alexander K W Elsholz, Kürsad Turgay, Stephan Michalik, Bernd Hessling, Katrin Gronau, Dan Oertel, Ulrike Mäder, Jörg Bernhardt, Dörte Becher, Michael Hecker, Ulf Gerth
Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis.
Proc Natl Acad Sci U S A: 2012, 109(19);7451-6
[PubMed:22517742]
[WorldCat.org]
[DOI]
(I p)
Pierre Nicolas, Ulrike Mäder, Etienne Dervyn, Tatiana Rochat, Aurélie Leduc, Nathalie Pigeonneau, Elena Bidnenko, Elodie Marchadier, Mark Hoebeke, Stéphane Aymerich, Dörte Becher, Paola Bisicchia, Eric Botella, Olivier Delumeau, Geoff Doherty, Emma L Denham, Mark J Fogg, Vincent Fromion, Anne Goelzer, Annette Hansen, Elisabeth Härtig, Colin R Harwood, Georg Homuth, Hanne Jarmer, Matthieu Jules, Edda Klipp, Ludovic Le Chat, François Lecointe, Peter Lewis, Wolfram Liebermeister, Anika March, Ruben A T Mars, Priyanka Nannapaneni, David Noone, Susanne Pohl, Bernd Rinn, Frank Rügheimer, Praveen K Sappa, Franck Samson, Marc Schaffer, Benno Schwikowski, Leif Steil, Jörg Stülke, Thomas Wiegert, Kevin M Devine, Anthony J Wilkinson, Jan Maarten van Dijl, Michael Hecker, Uwe Völker, Philippe Bessières, Philippe Noirot
Condition-dependent transcriptome reveals high-level regulatory architecture in Bacillus subtilis.
Science: 2012, 335(6072);1103-6
[PubMed:22383849]
[WorldCat.org]
[DOI]
(I p)
Jean-Christophe Meile, Ling Juan Wu, S Dusko Ehrlich, Jeff Errington, Philippe Noirot
Systematic localisation of proteins fused to the green fluorescent protein in Bacillus subtilis: identification of new proteins at the DNA replication factory.
Proteomics: 2006, 6(7);2135-46
[PubMed:16479537]
[WorldCat.org]
[DOI]
(P p)
Christoph Freiberg, Nina A Brunner, Guido Schiffer, Thomas Lampe, Jens Pohlmann, Michael Brands, Martin Raabe, Dieter Häbich, Karl Ziegelbauer
Identification and characterization of the first class of potent bacterial acetyl-CoA carboxylase inhibitors with antibacterial activity.
J Biol Chem: 2004, 279(25);26066-73
[PubMed:15066985]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Masaya Fujita, Shane T Jensen, Patrick Eichenberger, José E González-Pastor, Jun S Liu, Richard Losick
The Spo0A regulon of Bacillus subtilis.
Mol Microbiol: 2003, 50(5);1683-701
[PubMed:14651647]
[WorldCat.org]
[DOI]
(P p)
Hailong Zhang, Zhiru Yang, Yang Shen, Liang Tong
Crystal structure of the carboxyltransferase domain of acetyl-coenzyme A carboxylase.
Science: 2003, 299(5615);2064-7
[PubMed:12663926]
[WorldCat.org]
[DOI]
(I p)