Difference between revisions of "RbsD"
| Line 118: | Line 118: | ||
=References= | =References= | ||
| + | <pubmed>12738765 7921236, </pubmed> | ||
# Kim MS, Shin J, Lee W, Lee HS, Oh BH. (2003) Crystal structures of RbsD leading to the identification of cytoplasmic sugar-binding proteins with a novel folding architecture. ''J Biol Chem. '' '''Jul 25;278(30):''' 28173-80. [http://www.ncbi.nlm.nih.gov/sites/entrez/12738765 PubMed] | # Kim MS, Shin J, Lee W, Lee HS, Oh BH. (2003) Crystal structures of RbsD leading to the identification of cytoplasmic sugar-binding proteins with a novel folding architecture. ''J Biol Chem. '' '''Jul 25;278(30):''' 28173-80. [http://www.ncbi.nlm.nih.gov/sites/entrez/12738765 PubMed] | ||
# Woodson K, Devine KM. (1994) Analysis of a ribose transport operon from Bacillus subtilis. ''Microbiology. '' '''Aug;140 ( Pt 8):''' 1829-38. [http://www.ncbi.nlm.nih.gov/sites/entrez/7921236 PubMed] | # Woodson K, Devine KM. (1994) Analysis of a ribose transport operon from Bacillus subtilis. ''Microbiology. '' '''Aug;140 ( Pt 8):''' 1829-38. [http://www.ncbi.nlm.nih.gov/sites/entrez/7921236 PubMed] | ||
# Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | ||
Revision as of 21:14, 8 June 2009
- Description: ribose ABC transporter (membrane protein)
| Gene name | rbsD |
| Synonyms | |
| Essential | no |
| Product | ribose ABC transporter (membrane protein) |
| Function | ribose uptake |
| MW, pI | 14 kDa, 4.987 |
| Gene length, protein length | 393 bp, 131 aa |
| Immediate neighbours | rbsK, rbsA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 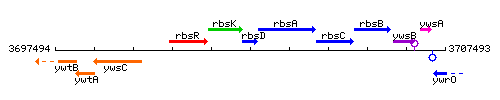
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU35930
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Beta-D-ribopyranose = beta-D-ribofuranose (according to Swiss-Prot)
- Protein family: RbsD subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure: 1OGD (complex with D-ribose)
- Swiss prot entry: P36946
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Min-Sung Kim, Joon Shin, Weontae Lee, Heung-Soo Lee, Byung-Ha Oh
Crystal structures of RbsD leading to the identification of cytoplasmic sugar-binding proteins with a novel folding architecture.
J Biol Chem: 2003, 278(30);28173-80
[PubMed:12738765]
[WorldCat.org]
[DOI]
(P p)
K Woodson, K M Devine
Analysis of a ribose transport operon from Bacillus subtilis.
Microbiology (Reading): 1994, 140 ( Pt 8);1829-38
[PubMed:7921236]
[WorldCat.org]
[DOI]
(P p)
- Kim MS, Shin J, Lee W, Lee HS, Oh BH. (2003) Crystal structures of RbsD leading to the identification of cytoplasmic sugar-binding proteins with a novel folding architecture. J Biol Chem. Jul 25;278(30): 28173-80. PubMed
- Woodson K, Devine KM. (1994) Analysis of a ribose transport operon from Bacillus subtilis. Microbiology. Aug;140 ( Pt 8): 1829-38. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed