Difference between revisions of "LicT"
| Line 22: | Line 22: | ||
|style="background:#FAF8CC;" align="center"|'''[http://subtiwiki.uni-goettingen.de/licT_nucleotide.txt Gene sequence (+200bp) ]''' | |style="background:#FAF8CC;" align="center"|'''[http://subtiwiki.uni-goettingen.de/licT_nucleotide.txt Gene sequence (+200bp) ]''' | ||
|style="background:#FAF8CC;" align="center"|'''[http://subtiwiki.uni-goettingen.de/licT_protein.txt Protein sequence]''' | |style="background:#FAF8CC;" align="center"|'''[http://subtiwiki.uni-goettingen.de/licT_protein.txt Protein sequence]''' | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;color:#FF0000" align="center" | '''Caution: The sequence for this gene in SubtiList contains errors | ||
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:licT_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:licT_context.gif]] | ||
Revision as of 12:26, 24 February 2009
- Description: transcriptional antiterminator of the bglPH operon
| Gene name | licT |
| Synonyms | |
| Essential | no |
| Product | transcriptional antiterminator (BglG family) |
| Function | required for substrate-dependent induction of bglPH |
| MW, pI | 32 kDa, 5.944 |
| Gene length, protein length | 831 bp, 277 aa |
| Immediate neighbours | bglS, yxiP |
| Gene sequence (+200bp) | Protein sequence |
| Caution: The sequence for this gene in SubtiList contains errors | |
Genetic context 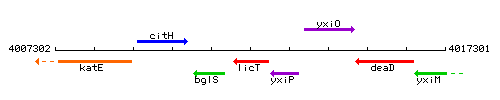
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
no expression of the bglP-bglH operon
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: binding to the mRNAs of bglS and the bglP-bglH operon, causes transcription antitermination (in presence of salicin and absence of glucose)
- Protein family: BglG family of antiterminators
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- Swiss prot entry:
- KEGG entry:
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Stephane Aymerich, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Josef Deutscher, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Michael Hecker, Greifswald, Germany Homepage
Your additional remarks
References
Original description
- Schnetz, K., Stülke, J., Gertz, S., Krüger, S., Krieg, M., Hecker, M. & Rak, B. (1996) LicT, a Bacillus subtilis transcriptional antiterminator protein of the BglG family. J. Bacteriol. 178: 1971-1979. PubMed
Control of LicT activity
- Lindner, C., Galinier, A., Hecker, M. & Deutscher, J. (1999) Regulation of the activity of the Bacillus subtilis antiterminator LicT by multiple PEP-dependent, enzyme I- and HPr-catalysed phosphorylation. Mol. Microbiol. 31, 995-1006 . PubMed
- Lindner, C., Hecker, M., Le Coq, D. & Deutscher, J. (2002) Bacillus subtilis mutant LicT antiterminators exhibiting enzyme I- and HPr-independent antitermination affect catabolite repression of the bglPH operon. J. Bacteriol. 184, 4819-4828 . PubMed
- Krüger, S., Gertz, S. & Hecker, M. Transcriptional analysis of bglPH expression in Bacillus subtilis: Evidence for two distinct pathways mediating carbon catabolite repression. J. Bacteriol. 178, 2637-2644 (1996). PubMed
- Tortosa, P., Declerck, N., Dutartre, H., Lindner, C., Deutscher, J., and Le Coq, D. (2001) Sites of positive and negative regulation in the Bacillus subtilis antiterminators LicT and SacY. Mol Microbiol 41: 1381-1393. PubMed
Structural analysis of LicT
- Declerck N, Dutartre H, Receveur V, Dubois V, Royer C, Aymerich S, van Tilbeurgh H (2001) Dimer stabilization upon activation of the transcriptional antiterminator LicT. J Mol Biol 314:671-681. PubMed
- Graille, M. et al. Activation of the LicT transcriptional antiterminator involves a domain swing/lock mechanism provoking massive structural changes. J. Biol. Chem. 280, 14780-14789 (2005). PubMed
- van Tilbeurgh H, Le Coq D, Declerck N (2001) Crystal structure of an activated form of the PTS regulation domain from the LicT transcriptional antiterminator. EMBO J 20:3789-3799. PubMed
- Déméné H, Ducat T, De Guillen K, Birck C, Aymerich S, Kochoyan M, Declerck N.(2008) Structural mechanism of signal transduction between the RNA-binding domain and the phosphotransferase system regulation domain of the LicT antiterminator.J. Biol. Chem. 283: 30838-30849. PubMed
LicT-RNA interaction
- Yang Y, Declerck N, Manival X, Aymerich S, Kochoyan M (2002) Solution structure of the LicT-RNA antitermination complex: CAT clamping RAT. EMBO J 21:1987-1997. PubMed
- Aymerich, S. and Steinmetz, M. (1992) Specificity determinants and structural features in the RNA target of the bacterial antiterminator proteins of the BglG/SacY family. Proc. Natl. Acad. Sci. USA 89, 10410-10414. PubMed
- Declerck, N., Vincent, F., Hoh, F., Aymerich, S. and van Tilbeurgh, H. (1999) RNA recognition by transcriptional antiterminators of the BglG/SacY family: functional and structural comparison of the CAT domains from SacY and LicT. J. Mol. Biol. 294, 389-402. PubMed
- Déméné H, Ducat T, De Guillen K, Birck C, Aymerich S, Kochoyan M, Declerck N.(2008) Structural mechanism of signal transduction between the RNA-binding domain and the phosphotransferase system regulation domain of the LicT antiterminator.J. Biol. Chem. 283: 30838-30849. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed