Difference between revisions of "SigM"
| Line 1: | Line 1: | ||
| − | * '''Description:''' [[RNA polymerase]] ECF-type [[sigma factor]] [[SigM]] <br/><br/> | + | * '''Description:''' [[RNA polymerase]] ECF-type [[sigma factor]] [[SigM]], responsible for intrinsic resistance against beta-lactam antibiotics <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 12: | Line 12: | ||
|style="background:#ABCDEF;" align="center"| '''Product''' || [[RNA polymerase]] ECF-type [[sigma factor]] [[SigM]] | |style="background:#ABCDEF;" align="center"| '''Product''' || [[RNA polymerase]] ECF-type [[sigma factor]] [[SigM]] | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || resistance against | + | |style="background:#ABCDEF;" align="center"|'''Function''' || responsible for intrinsic resistance against beta-lactam antibiotics |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://cellpublisher.gobics.de/subtinteract/startpage/start/ ''Subt''Interact]''': [http://cellpublisher.gobics.de/subtinteract/interactionList/2/SigM SigM] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://cellpublisher.gobics.de/subtinteract/startpage/start/ ''Subt''Interact]''': [http://cellpublisher.gobics.de/subtinteract/interactionList/2/SigM SigM] | ||
| Line 35: | Line 35: | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| − | |||
| − | |||
| − | |||
| − | |||
<br/><br/><br/> | <br/><br/><br/> | ||
| Line 44: | Line 40: | ||
{{SubtiWiki category|[[transcription]]}}, | {{SubtiWiki category|[[transcription]]}}, | ||
{{SubtiWiki category|[[sigma factors and their control]]}}, | {{SubtiWiki category|[[sigma factors and their control]]}}, | ||
| − | {{SubtiWiki category|[[cell envelope stress proteins (controlled by SigM, V, W, X, Y)]]}} | + | {{SubtiWiki category|[[cell envelope stress proteins (controlled by SigM, V, W, X, Y)]]}}, |
| + | {{SubtiWiki category|[[resistance against toxins/ antibiotics]]}} | ||
= This gene is a member of the following [[regulons]] = | = This gene is a member of the following [[regulons]] = | ||
| Line 58: | Line 55: | ||
* SigM is essential for growth and survival in nutrient broth (NB) containing 1.4 M NaCl {{PubMed|10216858}} | * SigM is essential for growth and survival in nutrient broth (NB) containing 1.4 M NaCl {{PubMed|10216858}} | ||
* ''sigM'' mutants form aberrantly shaped cells, which swell and lyse spontaneously during growth in NB medium containing increased levels (0.35-0.7 M) of a wide range of different salts {{PubMed|10216858}} | * ''sigM'' mutants form aberrantly shaped cells, which swell and lyse spontaneously during growth in NB medium containing increased levels (0.35-0.7 M) of a wide range of different salts {{PubMed|10216858}} | ||
| + | * increased sensitivity towards beta-lactam antibiotics {{PubMed|22211522}} | ||
=== Database entries === | === Database entries === | ||
| Line 147: | Line 145: | ||
<pubmed>18179421,17675383,17434969,</pubmed> | <pubmed>18179421,17675383,17434969,</pubmed> | ||
==Other original publications== | ==Other original publications== | ||
| − | <pubmed>22362028 18394148,14993308,12399481,12775685,12207695,18820022,15838020,19047346,14651641,16151214 ,19465659 19745567 10216858 20057163 | + | <pubmed>22362028 18394148,14993308,12399481,12775685,12207695,18820022,15838020,19047346,14651641,16151214 ,19465659 19745567 10216858 20057163 22211522 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 18:44, 21 July 2012
- Description: RNA polymerase ECF-type sigma factor SigM, responsible for intrinsic resistance against beta-lactam antibiotics
| Gene name | sigM |
| Synonyms | yhdM |
| Essential | no |
| Product | RNA polymerase ECF-type sigma factor SigM |
| Function | responsible for intrinsic resistance against beta-lactam antibiotics |
| Interactions involving this protein in SubtInteract: SigM | |
| MW, pI | 19 kDa, 6.77 |
| Gene length, protein length | 489 bp, 163 aa |
| Immediate neighbours | yhdL, yhdN |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 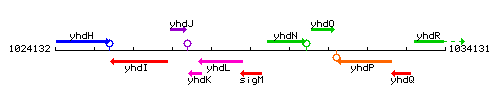
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed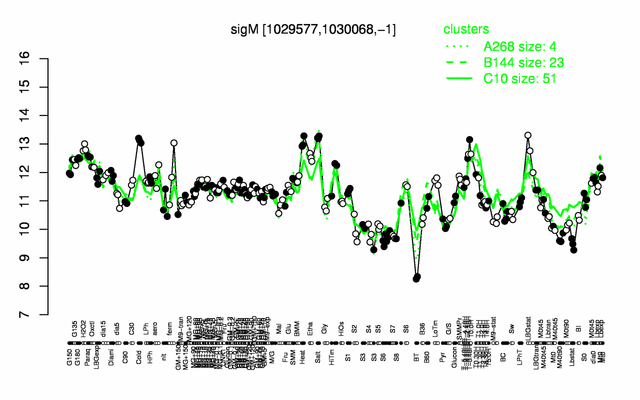
| |
Contents
Categories containing this gene/protein
transcription, sigma factors and their control, cell envelope stress proteins (controlled by SigM, V, W, X, Y), resistance against toxins/ antibiotics
This gene is a member of the following regulons
The SigM regulon
The gene
Basic information
- Locus tag: BSU09520
Phenotypes of a mutant
- SigM is essential for growth and survival in nutrient broth (NB) containing 1.4 M NaCl PubMed
- sigM mutants form aberrantly shaped cells, which swell and lyse spontaneously during growth in NB medium containing increased levels (0.35-0.7 M) of a wide range of different salts PubMed
- increased sensitivity towards beta-lactam antibiotics PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: sigma-70 factor family, ECF subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O07582
- KEGG entry: [3]
- E.C. number:
Additional information
- Expression of the SigM regulon in increased in ugtP mutants PubMed
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
John Helmann, Cornell University, USA Homepage
Your additional remarks
References
Additional publications: PubMed
The SigM regulon
Warawan Eiamphungporn, John D Helmann
The Bacillus subtilis sigma(M) regulon and its contribution to cell envelope stress responses.
Mol Microbiol: 2008, 67(4);830-48
[PubMed:18179421]
[WorldCat.org]
[DOI]
(P p)
Thorsten Mascher, Anna-Barbara Hachmann, John D Helmann
Regulatory overlap and functional redundancy among Bacillus subtilis extracytoplasmic function sigma factors.
J Bacteriol: 2007, 189(19);6919-27
[PubMed:17675383]
[WorldCat.org]
[DOI]
(P p)
Adrian J Jervis, Penny D Thackray, Chris W Houston, Malcolm J Horsburgh, Anne Moir
SigM-responsive genes of Bacillus subtilis and their promoters.
J Bacteriol: 2007, 189(12);4534-8
[PubMed:17434969]
[WorldCat.org]
[DOI]
(P p)
Other original publications