Difference between revisions of "AscR"
| Line 13: | Line 13: | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || regulation of sulphur metabolism | |style="background:#ABCDEF;" align="center"|'''Function''' || regulation of sulphur metabolism | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/riboflavin.html Riboflavin / FAD]''' | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 35 kDa, 6.228 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 35 kDa, 6.228 | ||
Revision as of 10:56, 16 February 2010
- Description: transcriptional activator of the ytmI-tcyJ-tcyK-tcyL-tcyM-tcyN-ytmO-ytnI-ytnJ-ribR-ytnL-ytnM operon
| Gene name | ytlI |
| Synonyms | |
| Essential | no |
| Product | transcriptional activator (LysR family) |
| Function | regulation of sulphur metabolism |
| Metabolic function and regulation of this protein in SubtiPathways: Riboflavin / FAD | |
| MW, pI | 35 kDa, 6.228 |
| Gene length, protein length | 924 bp, 308 aa |
| Immediate neighbours | ytmI, ytkL |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 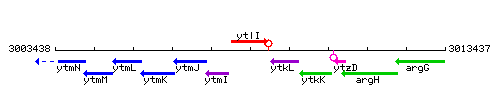
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU29400
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: LysR family
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- UniProt: O35038
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: ytlI PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Sergine Even, Pierre Burguière, Sandrine Auger, Olga Soutourina, Antoine Danchin, Isabelle Martin-Verstraete
Global control of cysteine metabolism by CymR in Bacillus subtilis.
J Bacteriol: 2006, 188(6);2184-97
[PubMed:16513748]
[WorldCat.org]
[DOI]
(P p)
Pierre Burguière, Juliette Fert, Isabelle Guillouard, Sandrine Auger, Antoine Danchin, Isabelle Martin-Verstraete
Regulation of the Bacillus subtilis ytmI operon, involved in sulfur metabolism.
J Bacteriol: 2005, 187(17);6019-30
[PubMed:16109943]
[WorldCat.org]
[DOI]
(P p)
Kyle N Erwin, Shunji Nakano, Peter Zuber
Sulfate-dependent repression of genes that function in organosulfur metabolism in Bacillus subtilis requires Spx.
J Bacteriol: 2005, 187(12);4042-9
[PubMed:15937167]
[WorldCat.org]
[DOI]
(P p)
Jean-Yves Coppée, Sandrine Auger, Evelyne Turlin, Agnieszka Sekowska, Jean-Pierre Le Caer, Valérie Labas, Valérie Vagner, Antoine Danchin, Isabelle Martin-Verstraete
Sulfur-limitation-regulated proteins in Bacillus subtilis: a two-dimensional gel electrophoresis study.
Microbiology (Reading): 2001, 147(Pt 6);1631-1640
[PubMed:11390694]
[WorldCat.org]
[DOI]
(P p)
Alia Lapidus, Nathalie Galleron, Alexei Sorokin, S Dusko Ehrlich
Sequencing and functional annotation of the Bacillus subtilis genes in the 200 kb rrnB-dnaB region.
Microbiology (Reading): 1997, 143 ( Pt 11);3431-3441
[PubMed:9387221]
[WorldCat.org]
[DOI]
(P p)