Difference between revisions of "CysC"
(→Expression and regulation) |
|||
| Line 92: | Line 92: | ||
* '''[[Sigma factor]]:''' [[SigA]] | * '''[[Sigma factor]]:''' [[SigA]] | ||
| − | * '''Regulation:''' repressed by casamino acids [http://www.ncbi.nlm.nih.gov/pubmed/12107147 PubMed] | + | * '''Regulation:''' |
| + | ** repressed by casamino acids [http://www.ncbi.nlm.nih.gov/pubmed/12107147 PubMed] | ||
| + | ** repressed in the presence of cysteine ([[CymR]]) | ||
| + | ** induced by methionine starvation ([[S-box]]) [http://www.ncbi.nlm.nih.gov/sites/entrez/10094622 PubMed] | ||
| − | * '''Regulatory mechanism:''' [[ | + | * '''Regulatory mechanism:''' |
| + | ** [[S-box]]: transcription termination/ antitermination, the [[S-box]] [[riboswitch]] binds S-adenosylmethionine resulting in termination [http://www.ncbi.nlm.nih.gov/sites/entrez/10094622 PubMed] | ||
| + | ** [[CymR]]: transcription repression | ||
* '''Additional information:''' | * '''Additional information:''' | ||
Revision as of 11:09, 7 June 2009
- Description: adenylyl-sulfate kinase
| Gene name | cysC |
| Synonyms | ylnC |
| Essential | no |
| Product | adenylyl-sulfate kinase |
| Function | sulfate reduction and activation |
| MW, pI | 22 kDa, 5.083 |
| Gene length, protein length | 591 bp, 197 aa |
| Immediate neighbours | sat, ylnD |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 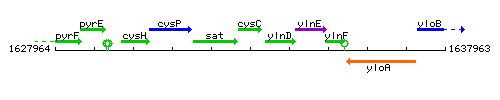
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU15600
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + adenylyl sulfate = ADP + 3'-phosphoadenylyl sulfate (according to Swiss-Prot)
- Protein family: APS kinase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- Swiss prot entry: O34577
- KEGG entry: [3]
- E.C. number: 2.7.1.25
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- S-box: transcription termination/ antitermination, the S-box riboswitch binds S-adenosylmethionine resulting in termination PubMed
- CymR: transcription repression
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Isabelle Martin-Verstraete, Institute Pasteur, Paris, France