Difference between revisions of "ComS"
| Line 57: | Line 57: | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| + | * overexpression of [[ComS]] results in expression of competence genes in 80% to 90% of the cells {{PubMed|8878039}} | ||
=== Database entries === | === Database entries === | ||
| Line 65: | Line 66: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
| Line 83: | Line 81: | ||
* '''Kinetic information:''' | * '''Kinetic information:''' | ||
| − | * '''Domains:''' | + | * '''[[Domains]]:''' |
* '''Modification:''' | * '''Modification:''' | ||
| − | * ''' | + | * '''[[Cofactors]]:''' |
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| Line 154: | Line 152: | ||
==Original Publications== | ==Original Publications== | ||
| − | <pubmed>17560370,10447896, 12642660,9000055, 10361283, 10447896, 11703662 1715856 16091051 20817675</pubmed> | + | <pubmed>17560370,10447896, 12642660,9000055, 10361283, 10447896, 11703662 1715856 16091051 20817675 8878039</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 11:33, 16 March 2014
- Description: antiadaptor protein, assembly link between regulatory components of the competence signal transduction pathway
| Gene name | comS |
| Synonyms | |
| Essential | no |
| Product | antiadaptor protein (anti-MecA) |
| Function | control of ComK degradation |
| Gene expression levels in SubtiExpress: comS | |
| Interactions involving this protein in SubtInteract: ComS | |
| MW, pI | 5 kDa, 9.145 |
| Gene length, protein length | 138 bp, 46 aa |
| Immediate neighbours | srfAB, srfAC |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 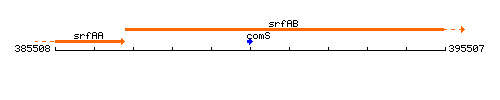
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed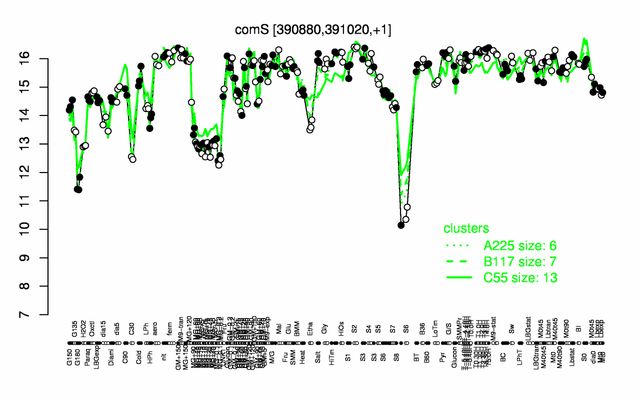
| |
Contents
Categories containing this gene/protein
genetic competence, short peptides
This gene is a member of the following regulons
Abh regulon, CodY regulon, ComA regulon, PerR regulon, Spx regulon
The gene
Basic information
- Locus tag: BSU03500
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P80355
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Janine Kirstein, Noël Molière, David A Dougan, Kürşad Turgay
Adapting the machine: adaptor proteins for Hsp100/Clp and AAA+ proteases.
Nat Rev Microbiol: 2009, 7(8);589-99
[PubMed:19609260]
[WorldCat.org]
[DOI]
(I p)
Original Publications
Onuma Chumsakul, Hiroki Takahashi, Taku Oshima, Takahiro Hishimoto, Shigehiko Kanaya, Naotake Ogasawara, Shu Ishikawa
Genome-wide binding profiles of the Bacillus subtilis transition state regulator AbrB and its homolog Abh reveals their interactive role in transcriptional regulation.
Nucleic Acids Res: 2011, 39(2);414-28
[PubMed:20817675]
[WorldCat.org]
[DOI]
(I p)
Peter Prepiak, David Dubnau
A peptide signal for adapter protein-mediated degradation by the AAA+ protease ClpCP.
Mol Cell: 2007, 26(5);639-47
[PubMed:17560370]
[WorldCat.org]
[DOI]
(P p)
Natalia Comella, Alan D Grossman
Conservation of genes and processes controlled by the quorum response in bacteria: characterization of genes controlled by the quorum-sensing transcription factor ComA in Bacillus subtilis.
Mol Microbiol: 2005, 57(4);1159-74
[PubMed:16091051]
[WorldCat.org]
[DOI]
(P p)
Shunji Nakano, Michiko M Nakano, Ying Zhang, Montira Leelakriangsak, Peter Zuber
A regulatory protein that interferes with activator-stimulated transcription in bacteria.
Proc Natl Acad Sci U S A: 2003, 100(7);4233-8
[PubMed:12642660]
[WorldCat.org]
[DOI]
(P p)
M M Nakano, F Hajarizadeh, Y Zhu, P Zuber
Loss-of-function mutations in yjbD result in ClpX- and ClpP-independent competence development of Bacillus subtilis.
Mol Microbiol: 2001, 42(2);383-94
[PubMed:11703662]
[WorldCat.org]
[DOI]
(P p)
M Persuh, K Turgay, I Mandic-Mulec, D Dubnau
The N- and C-terminal domains of MecA recognize different partners in the competence molecular switch.
Mol Microbiol: 1999, 33(4);886-94
[PubMed:10447896]
[WorldCat.org]
[DOI]
(P p)
M Ogura, L Liu, M Lacelle, M M Nakano, P Zuber
Mutational analysis of ComS: evidence for the interaction of ComS and MecA in the regulation of competence development in Bacillus subtilis.
Mol Microbiol: 1999, 32(4);799-812
[PubMed:10361283]
[WorldCat.org]
[DOI]
(P p)
K Turgay, L W Hamoen, G Venema, D Dubnau
Biochemical characterization of a molecular switch involving the heat shock protein ClpC, which controls the activity of ComK, the competence transcription factor of Bacillus subtilis.
Genes Dev: 1997, 11(1);119-28
[PubMed:9000055]
[WorldCat.org]
[DOI]
(P p)
J Hahn, A Luttinger, D Dubnau
Regulatory inputs for the synthesis of ComK, the competence transcription factor of Bacillus subtilis.
Mol Microbiol: 1996, 21(4);763-75
[PubMed:8878039]
[WorldCat.org]
[DOI]
(P p)
M M Nakano, L A Xia, P Zuber
Transcription initiation region of the srfA operon, which is controlled by the comP-comA signal transduction system in Bacillus subtilis.
J Bacteriol: 1991, 173(17);5487-93
[PubMed:1715856]
[WorldCat.org]
[DOI]
(P p)