Difference between revisions of "YkoV"
(→Reviews) |
(→Original publications) |
||
| Line 148: | Line 148: | ||
== Original publications == | == Original publications == | ||
| − | <pubmed>16497325, 11483577, 23691176 12215643 17293412</pubmed> | + | <pubmed>16497325, 11483577, 23691176 12215643 17293412 24123749 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 14:24, 21 October 2013
- Description: DNA-end-binding protein Ku, recruits LigD to DNA ends, confers dry-heat resistance to dormant spores PubMed
| Gene name | ykoV |
| Synonyms | |
| Essential | no |
| Product | DNA-end-binding protein Ku |
| Function | non-homologous end joining DNA repair, repair of gapped DNA substrates |
| Gene expression levels in SubtiExpress: ykoV | |
| Interactions involving this protein in SubtInteract: YkoV | |
| MW, pI | 34 kDa, 7.213 |
| Gene length, protein length | 933 bp, 311 aa |
| Immediate neighbours | ligD, dgcW |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 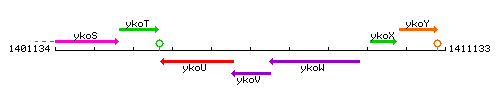
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed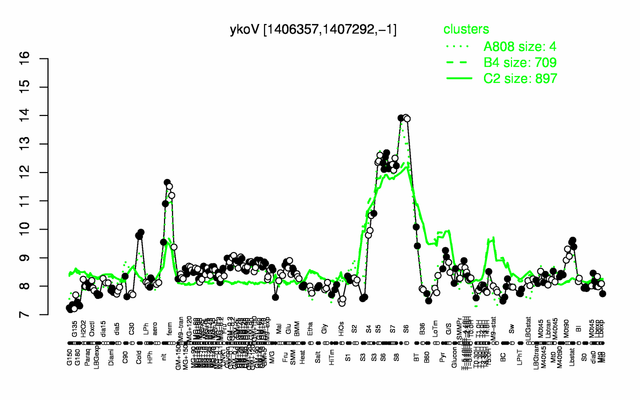
| |
Contents
Categories containing this gene/protein
DNA repair/ recombination, sporulation proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU13410
Phenotypes of a mutant
- sensitivity to ionizing radiation in the stationary phase PubMed
- sensitivity of spores to several DNA-damaging treatments known to cause double strand breaks, such as UV-ray, X-ray, ultrahigh vacuum and wet heat PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- associates with the nucleoid during germination PubMed
Database entries
- Structure:
- UniProt: O34859
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- BP141 (ykoV-ligD::kan) available in Fabian Commichau's lab
- BP142 (ykoW-ykoV-ligD::kan) available in Fabian Commichau's lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Justin S Lenhart, Jeremy W Schroeder, Brian W Walsh, Lyle A Simmons
DNA repair and genome maintenance in Bacillus subtilis.
Microbiol Mol Biol Rev: 2012, 76(3);530-64
[PubMed:22933559]
[WorldCat.org]
[DOI]
(I p)
Stewart Shuman, Michael S Glickman
Bacterial DNA repair by non-homologous end joining.
Nat Rev Microbiol: 2007, 5(11);852-61
[PubMed:17938628]
[WorldCat.org]
[DOI]
(I p)
A J Doherty, S P Jackson
DNA repair: how Ku makes ends meet.
Curr Biol: 2001, 11(22);R920-4
[PubMed:11719239]
[WorldCat.org]
[DOI]
(P p)
J M Jones, M Gellert, W Yang
A Ku bridge over broken DNA.
Structure: 2001, 9(10);881-4
[PubMed:11591342]
[WorldCat.org]
[DOI]
(P p)
Original publications
Ralf Moeller, Marina Raguse, Günther Reitz, Ryuichi Okayasu, Zuofeng Li, Stuart Klein, Peter Setlow, Wayne L Nicholson
Resistance of Bacillus subtilis spore DNA to lethal ionizing radiation damage relies primarily on spore core components and DNA repair, with minor effects of oxygen radical detoxification.
Appl Environ Microbiol: 2014, 80(1);104-9
[PubMed:24123749]
[WorldCat.org]
[DOI]
(I p)
Miguel de Vega
The minimal Bacillus subtilis nonhomologous end joining repair machinery.
PLoS One: 2013, 8(5);e64232
[PubMed:23691176]
[WorldCat.org]
[DOI]
(I e)
Ralf Moeller, Erko Stackebrandt, Günther Reitz, Thomas Berger, Petra Rettberg, Aidan J Doherty, Gerda Horneck, Wayne L Nicholson
Role of DNA repair by nonhomologous-end joining in Bacillus subtilis spore resistance to extreme dryness, mono- and polychromatic UV, and ionizing radiation.
J Bacteriol: 2007, 189(8);3306-11
[PubMed:17293412]
[WorldCat.org]
[DOI]
(P p)
Stephanie T Wang, Barbara Setlow, Erin M Conlon, Jessica L Lyon, Daisuke Imamura, Tsutomu Sato, Peter Setlow, Richard Losick, Patrick Eichenberger
The forespore line of gene expression in Bacillus subtilis.
J Mol Biol: 2006, 358(1);16-37
[PubMed:16497325]
[WorldCat.org]
[DOI]
(P p)
Geoffrey R Weller, Boris Kysela, Rajat Roy, Louise M Tonkin, Elizabeth Scanlan, Marina Della, Susanne Krogh Devine, Jonathan P Day, Adam Wilkinson, Fabrizio d'Adda di Fagagna, Kevin M Devine, Richard P Bowater, Penny A Jeggo, Stephen P Jackson, Aidan J Doherty
Identification of a DNA nonhomologous end-joining complex in bacteria.
Science: 2002, 297(5587);1686-9
[PubMed:12215643]
[WorldCat.org]
[DOI]
(I p)
L Aravind, E V Koonin
Prokaryotic homologs of the eukaryotic DNA-end-binding protein Ku, novel domains in the Ku protein and prediction of a prokaryotic double-strand break repair system.
Genome Res: 2001, 11(8);1365-74
[PubMed:11483577]
[WorldCat.org]
[DOI]
(P p)