Difference between revisions of "CitZ"
| Line 32: | Line 32: | ||
<br/><br/><br/><br/><br/><br/> | <br/><br/><br/><br/><br/><br/> | ||
| + | |||
| + | = [[Categories]] containing this gene/protein = | ||
| + | {{SubtiWiki category|[[carbon core metabolism]]}}, | ||
| + | {{SubtiWiki category|[[phosphoproteins]]}} | ||
| + | |||
| + | = This gene is a member of the following [[regulons]] = | ||
| + | {{SubtiWiki regulon|[[CcpA regulon]]}}, | ||
| + | {{SubtiWiki regulon|[[CcpC regulon]]}} | ||
=The gene= | =The gene= | ||
| Line 52: | Line 60: | ||
| − | + | ||
| − | |||
| − | |||
=The protein= | =The protein= | ||
Revision as of 22:20, 8 December 2010
- Description: citrate synthase
| Gene name | citZ |
| Synonyms | citA2 |
| Essential | no |
| Product | citrate synthase II |
| Function | TCA cycle |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 41 kDa, 5.451 |
| Gene length, protein length | 1116 bp, 372 aa |
| Immediate neighbours | icd, ytwI |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 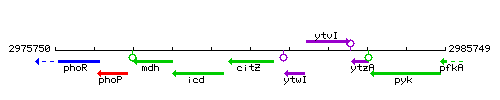
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
carbon core metabolism, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU29140
Phenotypes of a mutant
glutamate auxotrophy and a defect in sporulation PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Acetyl-CoA + H2O + oxaloacetate = citrate + CoA (according to Swiss-Prot)
- Protein family: citrate synthase family (according to Swiss-Prot)
Extended information on the protein
- Kinetic information: Michaelis-Menten (Random Sequential Reaction Mechanism) PubMed
- Domains:
- Modification: phosphorylation on Ser-284 PubMed
- Cofactor(s):
- Effectors of protein activity:
- Inhibited by acetyl-CoA, 2-oxoglutarate and NADH PubMed FEBS Letters
- Inhibited by citrate and CoA (competitively against acetyl-CoA and non-competitively against oxaloacetate) PubMed
- Inhibited by ATP competitively in B. subtilis strain 168 and HS 1A17 PubMed PubMed
- In B. subtilis strain HS 2A2, ATP inhibits a non-competitive fashion PubMed
- Activated by AMP PubMed
- Localization:
Database entries
- Structure: 2C6X
- UniProt: P39120
- KEGG entry: [3]
- E.C. number: 2.3.3.1
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
- The mRNA has a long 5' leader region. This may indicate RNA-based regulation PubMed
Biological materials
- Mutant: GP678 (erm), available in Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody: available in Linc Sonenshein lab
Labs working on this gene/protein
Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
Reviews
G Wiegand, S J Remington
Citrate synthase: structure, control, and mechanism.
Annu Rev Biophys Biophys Chem: 1986, 15;97-117
[PubMed:3013232]
[WorldCat.org]
[DOI]
(P p)
Original publications