Difference between revisions of "GapB"
(→References) |
(→References) |
||
| Line 120: | Line 120: | ||
=References= | =References= | ||
| − | # Meile et al. (2006) Systematic localisation of proteins fused to the green fluorescent protein in Bacillus subtilis: identification of new proteins at the DNA replication factory ''Proteomics'' '''6 | + | # Meile et al. (2006) Systematic localisation of proteins fused to the green fluorescent protein in ''Bacillus subtilis'': identification of new proteins at the DNA replication factory ''Proteomics'' '''6:''' 2135-2146. [http://www.ncbi.nlm.nih.gov/sites/entrez/16479537 PubMed] |
# Servant et al. (2005) CcpN (YqzB), a novel regulator for CcpA-independent catabolite repression of ''Bacillus subtilis'' gluconeogenic genes. Mol. Microbiol. 55: 1435-1451. [http://www.ncbi.nlm.nih.gov/sites/entrez/15720552 PubMed] | # Servant et al. (2005) CcpN (YqzB), a novel regulator for CcpA-independent catabolite repression of ''Bacillus subtilis'' gluconeogenic genes. Mol. Microbiol. 55: 1435-1451. [http://www.ncbi.nlm.nih.gov/sites/entrez/15720552 PubMed] | ||
# Tännler et al. (2008) CcpN controls central carbon fluxes in ''Bacillus subtilis''. J. Bacteriol. 190: 6178-6187. [http://www.ncbi.nlm.nih.gov/sites/entrez/18586936 PubMed] | # Tännler et al. (2008) CcpN controls central carbon fluxes in ''Bacillus subtilis''. J. Bacteriol. 190: 6178-6187. [http://www.ncbi.nlm.nih.gov/sites/entrez/18586936 PubMed] | ||
| − | # Thomaides, H. B., Davison, E. J., Burston, L., Johnson, H., Brown, D. R., Hunt, A. C., Errington, J., and Czaplewski, L. (2007) Essential bacterial functions encoded by gene pairs. J Bacteriol 189 | + | # Thomaides, H. B., Davison, E. J., Burston, L., Johnson, H., Brown, D. R., Hunt, A. C., Errington, J., and Czaplewski, L. (2007) Essential bacterial functions encoded by gene pairs. J Bacteriol 189: 591-602. [http://www.ncbi.nlm.nih.gov/sites/entrez/17114254 PubMed] |
Revision as of 19:31, 31 January 2009
- Description: glyceraldehyde-3-phosphate dehydrogenase, NADP-dependent, gluconeogenic enzyme
| Gene name | gapB |
| Synonyms | ppc |
| Essential | no |
| Product | glyceraldehyde-3-phosphate dehydrogenase 2 |
| Function | anabolic enzyme in gluconeogenesis |
| MW, pI | 37,3 kDa, 6.47 |
| Gene length, protein length | 1020 bp, 340 amino acids |
| Immediate neighbours | ytcD, speD |
| Gene sequence (+200bp) | Protein sequence |
Genetic context 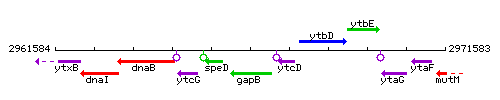
| |
Contents
The gene
Basic information
- Coordinates: 2966075 - 2967094
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: D-glyceraldehyde 3-phosphate + phosphate + NAD(P)(+) = 3-phospho-D-glyceroyl phosphate + NAD(P)H.
- Protein family: glyceraldehyde-3-phosphate dehydrogenase family
- Paralogous protein(s): GapA
Extended information on the protein
- Kinetic information:
- Domains:
- Nucleotid bindinge domain (12-13)
- 2x Glyceraldehyde 3-phosphate binding domain (151-153) & (210-211)
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cytoplasm
Database entries
- Structure:
- Swiss prot entry: [3]
- KEGG entry: [4]
- E.C. number: [5]
Additional information
Expression and regulation
- Operon:
- Regulatory mechanism: transcription repression
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- Antibody:
Labs working on this gene/protein
Stephane Aymerich, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Your additional remarks
References
- Meile et al. (2006) Systematic localisation of proteins fused to the green fluorescent protein in Bacillus subtilis: identification of new proteins at the DNA replication factory Proteomics 6: 2135-2146. PubMed
- Servant et al. (2005) CcpN (YqzB), a novel regulator for CcpA-independent catabolite repression of Bacillus subtilis gluconeogenic genes. Mol. Microbiol. 55: 1435-1451. PubMed
- Tännler et al. (2008) CcpN controls central carbon fluxes in Bacillus subtilis. J. Bacteriol. 190: 6178-6187. PubMed
- Thomaides, H. B., Davison, E. J., Burston, L., Johnson, H., Brown, D. R., Hunt, A. C., Errington, J., and Czaplewski, L. (2007) Essential bacterial functions encoded by gene pairs. J Bacteriol 189: 591-602. PubMed