Difference between revisions of "MinC"
| Line 125: | Line 125: | ||
* '''Regulation:''' | * '''Regulation:''' | ||
** expression is reduced in a [[SigV]] mutant {{PubMed|21926231}} | ** expression is reduced in a [[SigV]] mutant {{PubMed|21926231}} | ||
| + | ** constitutively expressed {{PubMed|23701187}} | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| Line 153: | Line 154: | ||
<pubmed> 19680248 19884039 </pubmed> | <pubmed> 19680248 19884039 </pubmed> | ||
==Original Publications== | ==Original Publications== | ||
| − | <pubmed>8459776,19019154,1400224,10411726,15317782, 20352045,18588879, 19141479 20566861 20711458 18179421 22457634 23577149 21926231</pubmed> | + | <pubmed>8459776,19019154,1400224,10411726,15317782, 20352045,18588879, 19141479 20566861 20711458 18179421 22457634 23577149 21926231 23701187</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 16:59, 27 May 2013
- Description: cell-division inhibitor (septum placement), destabilizes FtsZ-rings at new cell poles, part of the Min system (with DivIVA, MinD, MinJ), Noc and the Min system ensure the efficient utilization of the division site at midcell in by ensuring Z ring placement
| Gene name | minC |
| Synonyms | |
| Essential | no |
| Product | cell-division inhibitor |
| Function | septum placement |
| Gene expression levels in SubtiExpress: minC | |
| Interactions involving this protein in SubtInteract: MinC | |
| MW, pI | 24 kDa, 6.262 |
| Gene length, protein length | 678 bp, 226 aa |
| Immediate neighbours | minD, mreD |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 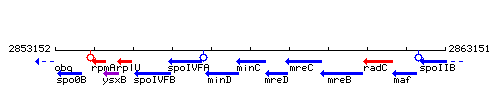
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed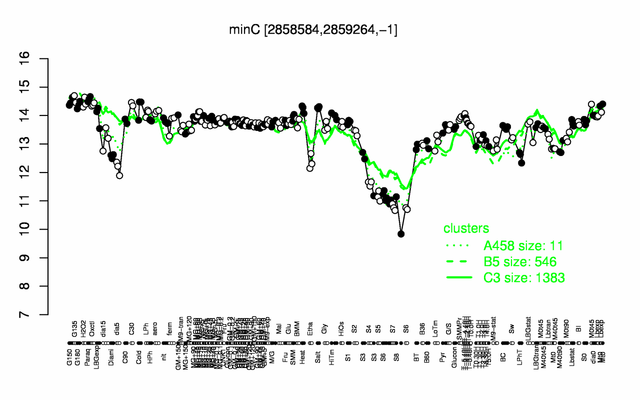
| |
Contents
Categories containing this gene/protein
cell division, cell envelope stress proteins (controlled by SigM, V, W, X, Y), membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU28000
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: minC family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: Q01463
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original Publications
Erik Nico Trip, Jan-Willem Veening, Eric J Stewart, Jeff Errington, Dirk-Jan Scheffers
Balanced transcription of cell division genes in Bacillus subtilis as revealed by single cell analysis.
Environ Microbiol: 2013, 15(12);3196-209
[PubMed:23701187]
[WorldCat.org]
[DOI]
(I p)
Valdir Blasios, Alexandre W Bisson-Filho, Patricia Castellen, Maria Luiza C Nogueira, Jefferson Bettini, Rodrigo V Portugal, Ana Carolina M Zeri, Frederico J Gueiros-Filho
Genetic and biochemical characterization of the MinC-FtsZ interaction in Bacillus subtilis.
PLoS One: 2013, 8(4);e60690
[PubMed:23577149]
[WorldCat.org]
[DOI]
(I p)
Christopher D A Rodrigues, Elizabeth J Harry
The Min system and nucleoid occlusion are not required for identifying the division site in Bacillus subtilis but ensure its efficient utilization.
PLoS Genet: 2012, 8(3);e1002561
[PubMed:22457634]
[WorldCat.org]
[DOI]
(I p)
Veronica Guariglia-Oropeza, John D Helmann
Bacillus subtilis σ(V) confers lysozyme resistance by activation of two cell wall modification pathways, peptidoglycan O-acetylation and D-alanylation of teichoic acids.
J Bacteriol: 2011, 193(22);6223-32
[PubMed:21926231]
[WorldCat.org]
[DOI]
(I p)
Inês Filipa Fernandes de Oliveira, Anabela de Sousa Borges, Viola Kooij, Jeremy Bartosiak-Jentys, Joen Luirink, Dirk-Jan Scheffers
Characterization of ftsZ mutations that render Bacillus subtilis resistant to MinC.
PLoS One: 2010, 5(8);e12048
[PubMed:20711458]
[WorldCat.org]
[DOI]
(I e)
Henrik Strahl, Leendert W Hamoen
Membrane potential is important for bacterial cell division.
Proc Natl Acad Sci U S A: 2010, 107(27);12281-6
[PubMed:20566861]
[WorldCat.org]
[DOI]
(I p)
Suey van Baarle, Marc Bramkamp
The MinCDJ system in Bacillus subtilis prevents minicell formation by promoting divisome disassembly.
PLoS One: 2010, 5(3);e9850
[PubMed:20352045]
[WorldCat.org]
[DOI]
(I e)
James A Gregory, Eric C Becker, Kit Pogliano
Bacillus subtilis MinC destabilizes FtsZ-rings at new cell poles and contributes to the timing of cell division.
Genes Dev: 2008, 22(24);3475-88
[PubMed:19141479]
[WorldCat.org]
[DOI]
(P p)
Marc Bramkamp, Robyn Emmins, Louise Weston, Catriona Donovan, Richard A Daniel, Jeff Errington
A novel component of the division-site selection system of Bacillus subtilis and a new mode of action for the division inhibitor MinCD.
Mol Microbiol: 2008, 70(6);1556-69
[PubMed:19019154]
[WorldCat.org]
[DOI]
(I p)
Dirk-Jan Scheffers
The effect of MinC on FtsZ polymerization is pH dependent and can be counteracted by ZapA.
FEBS Lett: 2008, 582(17);2601-8
[PubMed:18588879]
[WorldCat.org]
[DOI]
(P p)
Warawan Eiamphungporn, John D Helmann
The Bacillus subtilis sigma(M) regulon and its contribution to cell envelope stress responses.
Mol Microbiol: 2008, 67(4);830-48
[PubMed:18179421]
[WorldCat.org]
[DOI]
(P p)
David E Anderson, Frederico J Gueiros-Filho, Harold P Erickson
Assembly dynamics of FtsZ rings in Bacillus subtilis and Escherichia coli and effects of FtsZ-regulating proteins.
J Bacteriol: 2004, 186(17);5775-81
[PubMed:15317782]
[WorldCat.org]
[DOI]
(P p)
A L Marston, J Errington
Selection of the midcell division site in Bacillus subtilis through MinD-dependent polar localization and activation of MinC.
Mol Microbiol: 1999, 33(1);84-96
[PubMed:10411726]
[WorldCat.org]
[DOI]
(P p)
S Lee, C W Price
The minCD locus of Bacillus subtilis lacks the minE determinant that provides topological specificity to cell division.
Mol Microbiol: 1993, 7(4);601-10
[PubMed:8459776]
[WorldCat.org]
[DOI]
(P p)
P A Levin, P S Margolis, P Setlow, R Losick, D Sun
Identification of Bacillus subtilis genes for septum placement and shape determination.
J Bacteriol: 1992, 174(21);6717-28
[PubMed:1400224]
[WorldCat.org]
[DOI]
(P p)