Difference between revisions of "PdxK"
| Line 99: | Line 99: | ||
* '''Operon:''' ''pdxK'' {{PubMed|3100393}} | * '''Operon:''' ''pdxK'' {{PubMed|3100393}} | ||
| − | * '''[ | + | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=pdxK_3900963_3901778_1 pdxK] {{PubMed|22383849}} |
| + | |||
| + | * '''Sigma factor:''' | ||
* '''Regulation:''' | * '''Regulation:''' | ||
| Line 105: | Line 107: | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| − | * '''Additional information:''' subject to Clp-dependent proteolysis upon glucose starvation [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] | + | * '''Additional information:''' subject to Clp-dependent proteolysis upon glucose starvation [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] |
=Biological materials = | =Biological materials = | ||
Revision as of 08:56, 17 April 2012
- Description: pyridoxine, pyridoxal, and pyridoxamine kinase
| Gene name | pdxK |
| Synonyms | ywdB, ipa-52r, thiD |
| Essential | no |
| Product | pyridoxine, pyridoxal, and pyridoxamine kinase |
| Function | biosynthesis of pyridoxal phosphate |
| MW, pI | 28 kDa, 4.922 |
| Gene length, protein length | 813 bp, 271 aa |
| Immediate neighbours | ywdC, ywdA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 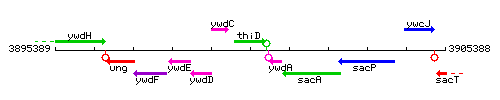
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU38020
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + 4-amino-2-methyl-5-phosphomethylpyrimidine = ADP + 4-amino-2-methyl-5-diphosphomethylpyrimidine (according to Swiss-Prot)
- Protein family: thiD family (according to Swiss-Prot)
- Paralogous protein(s): ThiD
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure: 2IB5
- UniProt: P39610
- KEGG entry: [3]
- E.C. number: 2.7.4.7
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Operon: pdxK PubMed
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Joseph A Newman, Sanjan K Das, Svetlana E Sedelnikova, David W Rice
Cloning, purification and preliminary crystallographic analysis of a putative pyridoxal kinase from Bacillus subtilis.
Acta Crystallogr Sect F Struct Biol Cryst Commun: 2006, 62(Pt 10);1006-9
[PubMed:17012797]
[WorldCat.org]
[DOI]
(I p)
Joo-Heon Park, Kristin Burns, Cynthia Kinsland, Tadhg P Begley
Characterization of two kinases involved in thiamine pyrophosphate and pyridoxal phosphate biosynthesis in Bacillus subtilis: 4-amino-5-hydroxymethyl-2methylpyrimidine kinase and pyridoxal kinase.
J Bacteriol: 2004, 186(5);1571-3
[PubMed:14973012]
[WorldCat.org]
[DOI]
(P p)
A Fouet, A Klier, G Rapoport
Nucleotide sequence of the sucrase gene of Bacillus subtilis.
Gene: 1986, 45(2);221-5
[PubMed:3100393]
[WorldCat.org]
[DOI]
(P p)