Difference between revisions of "SerA"
(→Extended information on the protein) |
|||
| Line 45: | Line 45: | ||
* '''Locus tag:''' BSU23070 | * '''Locus tag:''' BSU23070 | ||
| + | |||
| + | [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=serA_2411086_2412663_1 Expression] | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
Revision as of 12:47, 25 January 2012
- Description: phosphoglycerate dehydrogenase
| Gene name | serA |
| Synonyms | |
| Essential | no |
| Product | phosphoglycerate dehydrogenase |
| Function | biosynthesis of serine |
| Metabolic function and regulation of this protein in SubtiPathways: Ala, Gly, Ser | |
| MW, pI | 56 kDa, 5.617 |
| Gene length, protein length | 1575 bp, 525 aa |
| Immediate neighbours | ypzE, aroC |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 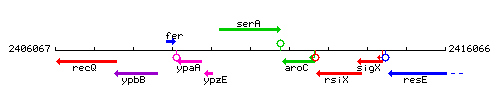
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU23070
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 3-phospho-D-glycerate + NAD+ = 3-phosphonooxypyruvate + NADH (according to Swiss-Prot)
- Protein family: D-isomer specific 2-hydroxyacid dehydrogenase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: membrane PubMed
Database entries
- UniProt: P35136
- KEGG entry: [3]
- E.C. number: 1.1.1.95
Additional information
Expression and regulation
- Operon: serA PubMed
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Bui Khanh Chi, Katrin Gronau, Ulrike Mäder, Bernd Hessling, Dörte Becher, Haike Antelmann
S-bacillithiolation protects against hypochlorite stress in Bacillus subtilis as revealed by transcriptomics and redox proteomics.
Mol Cell Proteomics: 2011, 10(11);M111.009506
[PubMed:21749987]
[WorldCat.org]
[DOI]
(I p)
Hannes Hahne, Susanne Wolff, Michael Hecker, Dörte Becher
From complementarity to comprehensiveness--targeting the membrane proteome of growing Bacillus subtilis by divergent approaches.
Proteomics: 2008, 8(19);4123-36
[PubMed:18763711]
[WorldCat.org]
[DOI]
(I p)
Falko Hochgräfe, Jörg Mostertz, Dierk-Christoph Pöther, Dörte Becher, John D Helmann, Michael Hecker
S-cysteinylation is a general mechanism for thiol protection of Bacillus subtilis proteins after oxidative stress.
J Biol Chem: 2007, 282(36);25981-5
[PubMed:17611193]
[WorldCat.org]
[DOI]
(P p)
James R Thompson, Jessica K Bell, Judy Bratt, Gregory A Grant, Leonard J Banaszak
Vmax regulation through domain and subunit changes. The active form of phosphoglycerate dehydrogenase.
Biochemistry: 2005, 44(15);5763-73
[PubMed:15823035]
[WorldCat.org]
[DOI]
(P p)
V Azevedo, A Sorokin, S D Ehrlich, P Serror
The transcriptional organization of the Bacillus subtilis 168 chromosome region between the spoVAF and serA genetic loci.
Mol Microbiol: 1993, 10(2);397-405
[PubMed:7934830]
[WorldCat.org]
[DOI]
(P p)